Just a few days ago the preprint of a new study on the prehistory of the North Caucasus was released (Wang et al. 2018). This is a major paper for understanding the West Eurasian prehistory, bringing (possibly) a few long awaited answers, but also several new interesting questions.
Archaeogenetic studies have described the formation of Eurasian ‘steppe ancestry’ as a mixture of Eastern and Caucasus hunter-gatherers. However, it remains unclear when and where this ancestry arose and whether it was related to a horizon of cultural innovations in the 4th millennium BCE that subsequently facilitated the advance of pastoral societies likely linked to the dispersal of Indo-European languages.
Indeed, the exact origin and genesis of the so called ‘steppe ancestry’ (AKA Yamnaya ancestry) has been quite a mystery. We knew that this type of ancestry could be described as a mixture of Eastern Hunter Gatherers (EHG) from the Forest Steppe and Steppe river banks of Eastern Europe and Caucasus Hunter Gatherers (CHG) from the South Caucasus. But somehow this was always problematic, due to the geographical separation of these groups, the specific mtDNA and Y-Chromosome profile of the putative resulting group (suggesting a strong sex biased admixture) and the fact that to a degree, while the model worked, it was never really as good as you would expect if you had the right populations.
We observe a genetic separation between the groups of the Caucasus and those of the adjacent steppe. The Caucasus groups are genetically similar to contemporaneous populations south of it, suggesting that – unlike today – the Caucasus acted as a bridge rather than an insurmountable barrier to human movement.
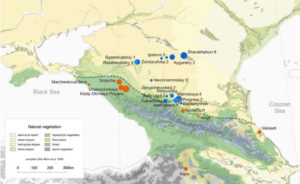
Indeed, it seems that for the Neolithic Farmers, the Great Caucasus wasn’t such a big barrier (though it seems to have slowed down the penetration of the Neolithic for quite a while), but the steppe itself does seem to have acted as a clear barrier for them, as can be seen by the dotted line in the map from the preprint:
As can be seen, the line dividing these two populations (orange and blue dots respectively) is in the border between the northern slopes of the Great Caucasus and the steppe in the North Caucasus area. This means that while farmers moved across the mountain range, they never moved beyond it, probably because the steppe was just not suitable for their subsistence economy (based not just on animal husbandry, but also on crops).
The paper focuses to a large degree on the differences between the two populations, and does not go much beyond that in terms of the consequences from these findings. So here I’ll focus a bit more on 3 samples labelled as “Eneolithic steppe” that seem to be key for understanding the genesis of the steppe Early Bronze Age populations and by extension the Bronze Age of most of West Eurasia:
- PG2001. Progress 2 site (see map above). 4336-4178 calBCE (charcoal), 4991-4834 calBCE (bone). mtDNA I3a, Y-DNA R1b1.
- PG2004. Progress 2 site. 4233-4047 calBCE. mtDNA H2, Y-DNA R1b1.
- VJ1001. Vonyuchka 1 site. 4332-4238 calBCE. mtDNA T2a1b. Female.
It’s somehow unfortunate that even if both males seem to have good coverage, their Y haplogroups could be determined in greater depth to see if they belong to R1b-M269. Maybe someone will try to test them further once the samples are publicly available. Their mtDNA also seems to be typical of steppe populations. Leaving aside the very generic H2, for the other two ones several matches can be found in CWC, Unetice, Poltavka and Andronovo (but none in ancient West Asia thus far).
Autosomally this population is very close to Yamnaya. For example, in Sup. table 13, Yamnaya_Samara is modelled as 88.9% Eneolithic_steppe, 5.2% Anatolia_N and 5.9% WHG (though I suspect that with something like Ukraine_N instead of WHG it might be a better fit). This would make this Eneolithic_Steppe population at roughly a 50/50 mix of EHG and CHG. The question now is if this is what actually happened in the area during the Eneolithic. And for me, the answer looks like a clear no.
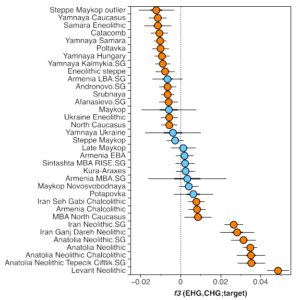
First, we do have contemporary samples from the Eneolithic North Caucasus foothills (I2055, I2056, I1722, c. 4500 calBCE), and just a quick look at the PCA on the main paper (Fig. 2D) tells us that they are way too western to be a suitable source. That kind of mix would never work for those Eneolithic_steppe samples. And then we have the f3 admixture tests checking for CHG-EHG admixture and the signal in the Eneolithic_steppe samples is weaker than in the much later Yamnaya groups (Sup. Fig. 3):
This probably means that the population is old in the area, probably Mesolithic, and Yamnaya got some extra EHG-related admixture that increases the signal. It also has to be kept in mind that before the invention of wheeled vehicles (and probably the domestication of the horse), no one was really living in the open steppe, where survival would have been very difficult. Mobility was limited to the river routes, and the Forest steppe was very distant from the Caucasus mountains to have any close contacts.
So what does this all mean? Well, I think a few interesting points that I’ll just briefly outline to not extend this post too much:
- Firstly, it does away with the idea that a large influx from south of the Caucasus entered the steppe to form the Yamnaya population. As far as I can tell, hardly any gene flow there (Yamnaya got just a bit – say 10%- probably from Ukraine Eneolithic groups, which in turn would be a mix of Ukraine_N and Tripolye (?) farmers. This weakens the idea that if this pre-Yamnaya population spoke PIE, the language came from the south (though the cultural impact from the south was still very big, so it’s not only genes that are involved in language transmission. A more nuanced and multidisciplinary approach would be needed in this case).
- Secondly, it also questions the validity of the “Fourth strand” model for the genetic makeup of Europe, in which modern Europeans would derive most of their ancestry from 4 early Holocene groups: WHG, Anatolian_Neolithic, EHG and CHG. Yes, I realise that ultimately, this North Caucasus steppe population is intermediate between EHG and CHG, but it could be so just in the same way as EHG is intermediate between ANE and WHG, or CHG themselves are intermediate between a putative Basal Eurasian population and an ancient West Eurasian one. So it all depends on how far one goes, but the “Mesolithic” model now looks like Europeans derive most of their ancestry from just 3 main populations instead of 4.
- Related with all the above, and again referring to the linguistic implications, it also puts into question the model where IE languages and Uralic languages have a genetic relationship deep into prehistory. If the language of this new found population was ancestral to IE (by the time of the samples it actually should be PIE itself), there’s no evidence of early contacts with Uralic, unless one places Uralic maybe in the Lower Volga or Lower Don? More likely, the contacts are much later, between an archaic Indo-Iranian language spoken in the steppe from around 1500 BCE and proto-Uralic?
There are many, many more things to debate in this very interesting paper (Maykop itself, the arrival of Iranian Farmers at around 3500 BCE in the Novosvobodnaya phase/culture, the arrival of Botai-related ancestry to the North Caucasus steppe c. 3200 BCE, etc… but I’ll leave that for either the comments section of for a later post.
P.S: I did want to make a small comment about one conclusion in the paper regarding the modern difference between populations from the North Caucasus and the South Caucasus:
First, sometime after the Bronze Age present-day North Caucasian populations must have received additional gene-flow from populations north of the mountain range that separates them from southern Caucasians, who largely retained the Bronze Age ancestry profile.
I don’t know, but it seems to me that the very “Yamnaya-like” profile seen in modern North Caucasian populations derives in great part from their putative Mesolithic ancestors who were “right there”, not in some distant northern location. So the distinction between north and south seems easy to explain by that ancient difference without the need to refer to historical events (which obviously happened, but don’t necessarily explain the genetic difference. More about this in a future post).

There really is not much information about those Eneolithic steppe sites, but something caught my attention: there are several references this paper that I couldn’t find online:
Gresky J., Berezina N.Ya. Two cases of trepanation in Eneolithic burials from Progress 2 and Vonjuchka 1
So those seem to be the exact samples I mentioned in the post. And I remembered an article from a couple of years ago that talked about this practice in Southern Russia:
New cases of trepanations from the 5th to 3rd millennia BC in Southern Russia in the context of previous research: Possible evidence for a ritually motivated tradition of cranial surgery?
So apparently this “tradition” went on from the 5th mill. that these samples are from all the way to the 3rd mill. Which means that there was a continuity between the Eneolithic and Bronze age populations in some way (apart from genetic continuity).
This was a much needed paper. It is fascinating to see structure in the north Caucasus region (sensu latu) – the Eneolithic steppe which might be a precursor to later (eastern) Yamnaya, ‘exotic’ Siberian ancestry (Q1a) individuals, and southern migrants in the foothills. It curious to see a genetic barrier between the Steppe Eneolithic & Majkop, despite the sharing of some key cultural elements.
Also, it would be great to clarify further the mesolithic – Neolithic – eneolithic sequence of the region to help understand the genetics.
I also think that these results are very significant. It seems that we have all the necessary ingredients for a steppe expansion from the Caucasus steppe:
right time frame
high level of metallurgy from Maikop
kurgans
horse culture
Yamnaya mixture of EHG and CHG
yDNA R1b-M269
I took a look at my mtDNA database and found the following interesting matches for I3a, H2 and T2a1b:
I3a:
Poltavka Lopatino II Samara I0440/SVP53 I3a,
BB Germany-BAV Alburg-Lerchenhaid I3602 I3a
BB Hungary Budakalász I3529/GEN86 I3a,
Unetice EBA Esperstedt Germany I0114 + I0117 I3a,
Unetice EBA Benzingerode-Heimburg Germany BZH18 I3a,
North Wales Early Bronze Age Great Orme Mines I1775/GOM245
Scythian Iron Age Rostov-on-Don Russia RD14 + RD15 I3
H2:
Dnieper Yamnaya Kirovograd Sugokleya Ukraine SUG7 + SUG8 H2
Manych Catacomb Temrta III Ukraine TEM2 H2
Lithuania LBA Turlojiske RISE598 H2a2
Globular Amphora Culture H2a2
Sarmatian Sarmatian6 Chebotarev V Rostov region Russia DA144 H2
T2a1b:
Donau Yamnaya Golyamata Mogila Bulgaria POP1 T2a1b1a
Corded Ware Esperstedt Germany I0106/ESP26 T2a1b1
Bell Beaker Benzingerode Heimburg Germany BZH5 T2a1b
Scotland Early Bronze Age Orkney I2981 T2a1b1a,
EBA Scotland Covesea Cave Moray I3132 2000 BC T2a1b1a,
Early Bronze Age Scotland Stenchme Orkney I2981 1750 BCE T2a1b1a
Andronovo Solenoozernaïa IV Russia S11 T2a1b1
Udegram IA Khyber-Pakhtunkwa Swat Babozai tahsil Pakistan SPGT S8191.E1.L1 T2a1b
Udegram IA Khyber-Pakhtunkwa Swat Babozai tahsil Pakistan SPGT I1799 T2a1b
In general, the mtDNA data of this paper is very interesting. Some mtDNA haplotypes fit very well the IE trail:
Late North Caucasus Kabardinka kurgan KBD001 2000 BC mtDNA I4a, yDNA R1b-L23
BB Central Europe Radovesice Czech I7286 I4a,
BB Central Europe Prague Czech I4946 I4a
BB Central Europe Augsburg Bavaria I5519 I4a
England CA/Early Bronze Age Wiltshire I2457 I4a
EBA Czech Moravská Nová Ves I5042/RISE584 I4a
Baltia LN Lithuania Spiginas2 c. 1900 BC I4a,
Iron Age Wielbark Kow-OVIA Poland PCA0040 100AD I4a
Some southern mtDNAs are interestingly shared between yDNA J and R1b rich populations:
Eneolithic Caucasus Unakozovskaya Russia I2055 R1a (yDNA J)
Eneolithic Caucasus Unakozovskaya Russia I2056 R1a (yDNA J2a)
Eneolithic Caucasus Unakozovskaya Russia I1722 R1a
Maykop Novosvobodnaya Klady kurgan I6268 R1a
Kura-Araxes Kaps Armenia ARM001.A0101 R1a1
Yamnaya Karagash EBA Kazakhstan RISE786 R1a1a (yDNA R1b-Z2108)
EBA Armenia Kura Araxes Kaps arm2 R1a1
North Caucasus Rasshevatskiy 1 kurgan Russia RK1003.C0101 R1a1a
North Caucasus Marinskaya 5 kurgan MK5009.A0101 R1a1a (yDNA R1b-L23)
EIA Armenia Lchashen-Metsamor Kanagegh arm23 R1a1a
EBA Germany Haunstetten Unterer Talweg Massy153 R1a1a
BB Central Europe Brandýsek Czech I7250 R1a1a
BB Britain Over Narrows I2454/OVE08 R1a1a
(BB means Bell Beaker )
R1a is fairly rare in modern Europeans, but it is present in Russians, Swedes and Finns and in IE speaking Armenians and Iranians; in India it is an IE caste haplogroup.
North Caucasus Goryachevodskiy 2 kurgan GW1001.A0101 2867-2581 BC mtDNA U2e1b (yDNA R1b1a2a2-Z2103)
U2e1b is found in the following modern groups:
Anatolia; Poland, Ukraine, Russia, Belarus, Denmark, Finland U2e1b1; Tula Russia U2e1b2 + Italy, Germany, UK; India U2e1b (IE upper caste, some Dravidians); Siberia Bargut U2e1b3
These haplotypes which look like radiating from the Bronze Age Caucasus are usually not high frequency haplogroups but the most important thing is the trail that they leave behind.
@Kristiina
That’s quite an impressive list of mtDNA samples across West Eurasia that match with those from the North Caucasus. Very interesting the case of R1a that kind of links the populations at both sides of the line and expanded with them.
@Robert
Now I remembered that there seem to be some UP samples from the North Caucasus coming. Someone mentioned in a comment at Eurogenes a while back that they were from around 20-23 Kya (IIRC) and already had most of the mutations for light pigmentation.
Those might be too early to have Basal Eurasian admixture, but there’s still plenty of time since the Satsurblia sample (c. 13 Kya?) for it to have arrived to the North Caucasus long before the arrival of the Neolithic.
Quite surprising that the area is archaeologically not very well researched. It might turn out to be quite crucial for the genetic history of West Eurasia.
For me, the results in Supp. figures 13 and 14 are quite interesting in that Sintashta and Andronovo models essentially fail relative to the rest. Is this merely indicative of them and/or their precursor populations being highly drifted? Or does this suggest some fundamentally different dynamics in their genesis?
I wish they’d included test CWC populations from Poland, Germany and the Baltic for comparison, btw.
What do you think about the role of the once wet Manych-Kerch spillway in the formation of the earliest basal-rich(Zarzianate?) population pool north of the Caucasus? This is pure speculation at this point, but my guess is that the spillway could have served a dual function: 1)acted as a barrier b/ween Crown Eurasians north of it 2)provided a lush, riparian environment(the importance of which you also surmise) conducive to a local maximum in population density north of the caucasus.
Note that Caucasus and Eneolithic Steppe demes(as defined by paper) take different CHG-related nodes. While they both take smth to the exclusion of the EEF-related node, Eneo Steppe’s node is an outgroup relative to CHG and the inferred CHG-related node for Caucasus populations.
We’re looking at 15-12KYA for the spillway.
@Anthro Survey
For Supp. Table 13 I would think that the lack of any sort of EHG (or at least Ukraine_N) could be making some models fail, since CWC and derivatives seem to have extra EHG on top of Yamnaya (and the Eneolithic _Steppe has probably less EHG than Yamnaya proper).
But then in Supp. Table 14 with the classical 4 “strands”, several models fail too. Not sure why. Maybe EHG + CHG is not good enough to represent Yamnaya? No idea, something to look at when the samples are released.
As for the Manych-Kerch spillway, it’s an interesting idea, but my knowledge of the specifics of the archaeology involved there are very basic. Probably worth a closer look from my part.
@Alberto
Yes, exactly–I had the same reasoning re/the Table13 model being too rigid wrt EHG input, but, on closer inspection, the EHG:CHG ratio doesn’t vary dramatically between “classic Yamna” and Andronovo/Sintashta.
The Yamna models mostly do fine in Table14(and 13). Only Steppe_MLBA struggles there.
Yes, that’s true. Difficult to say why that model isn’t working for steppe_MLBA. Maybe 4 distant sources combine for a sub-par model, or maybe we’re missing something.
Anyway, moving onto other things, finally we have the long awaited Maykop DNA. A first look reveals that certainly this was a new population in the area. The previous Eneolithic_Caucasus population looks more or less like a straight mix of CHG and Anatolia_N, in some 70/30 ratio approx. It should represent the South Caucasus Neolithic, I suppose.
Maykop sees a shift towards Europe_N. I wouldn’t say it necessarily means direct input from Europe, but it’s likely that there is some. It seems quite related to the Armenia_ChL samples that we already know. These in turn are quite interesting, since they really have this European shift combined with a North Iran/SC Asian one. Maykop shares Y-DNA L too.
And then we have Novosvobodnaya as a related, but differentiated culture. It has a good amount of Iran_ChL admixture, together with the Iranian expertise in arsenical bronze working. It also brings the Y-DNA J2a1, already found in Mycenaean, Hittite and Swat Valley samples.
It mirrors quite accurately the shifts in the South Caucasus itself:
Caucasus_Eneolithic <=> Shulaveri-Shomu (?)
Maykop <=> Armenia_ChL
Novosvobodnaya <=> Kura Araxes
Still the main question is the cultural genesis of Maykop/Armenia_ChL. Is the cultural impulse coming from the remains of Varna? (To find a precedent in Maykop’s “Royal” kurgan). Is it a synthesis of both East and West? Or are the Eastern influences just a later layer in the Novosvobodnaya culture?
I don’t understand the conclusions. Can you state the premises more clearly in future posts?
The fact that groups with ‘Steppe Eneolithic’-like genetic profile could have existed in Europe during the Mesolithic, doesn’t mean that there wasn’t and infux from the ‘south’.
The ‘samples from the Eneolithic North Caucasus foothills’ are from one region in Adygea. Groups with different genetic profiles could have existed close to them and even south of them, around 4500 BCE and earlier.
@ Anthro
An epi-paleolithic entity known as the Imeratian or Caucasian ‘Epigravettian’ existed after 20 to c. 10 ky cal BP from south to NW Caucasus, and a find spot or two also on the lower Don. This could explain the haplogroup J1 making its way to Karelia & Popova, and the (?) long presence of CHG in the piedmont steppe.
As mentioned above, however, archaeology has to clarify what exactly happened between the early Mesolithic and the Meshoko & ‘Eneolithic steppe’ occupations.
Perhaps the north Caucasus (& Crimea) were transiently occupied – at certain periods- when south to north movements occurred, before the commencement of permanent settlement after 4500 BC.
@ Alberto
” Someone mentioned in a comment at Eurogenes a while back that they were from around 20-23 Kya (IIRC) ”
Yes i remember, but then it disappeared 🙂
Do you remember the site they said ?
I can’t think of any such material from 23 kya (? maybe new find). There is the Satanay skull, but that is just a skull cap (therefore probably not viable for aDNA extraction) and from the turn of the Mesolithic.
@Apóstolos
My premise is that we have the samples of the Neolithic people who moved to the North Caucasus from the South Caucasus in the early 5th mill. BCE, and those are not a plausible match for admixture into the Eneolithic_steppe samples.
So definitely there was an influx from the south, but this was pre-Neolithic. Also given the low signal of admixture in the Eneolithic_steppe samples, I would think that whatever admixture happened it clearly predates these samples.
Of course the details of what and when it happened are unknown at this point. Could have been something from around 6000 BCE or around 15000 BCE, or anything else. There is no evidence that it was after around 6000 BCE, though only older samples will clarify that.
Anyway it’s only genetics. The Neolithic influx from the south, and then the Chalcolithic one, did have a big cultural impact on the steppe groups, that’s undeniable. But the gene flow at those stages seems to have been very limited.
@Robert
I found that old comment. No idea how reliable the source is, and it doesn’t mention any specific site, just the NW Caucasus:
http://eurogenes.blogspot.com/2017/01/higher-caste-lighter-skin.html?showComment=1483820791094#c516567292566366175
Alberto,
Nice to see your own blog. I wish you all the best and I hope this lasts long.
I have a somewhat radical hypothesis for the appearance of CHG/Iran_N admixture without an accompanying ANF admixture on the Eneolithic steppe and steppe Maykop.
As we know, the Anatolian_N (ANF) admixture in Iran is already evident as early as 5500 BCE and it already reached the North Caucasus by 4500 BCE. Therefore it would require a very early migration before 6000 BCE via the Caucasus for the CHG/Iran_N to reach unadmixed on the steppe.
So could it be that the Iran_N/CHG admixture in Eneolithic steppe & steppe Maykop is coming from across the Caspian from Central Asia. We already have several samples from there which are purely mixtures of Iran_N & WSHG/EHG such as Namazga, Sarazm, Dali_EBA.
It would make quite a bit of sense then since we also see the East Asian admixture in steppe Maykop.
Hi Jaydeep. Thanks! And welcome.
I don’t think your idea about the admixture in the Eneolithic steppe is something radical. I’ve entertained that idea myself since long ago, but lack of data was always a problem. Now that we have a much better sampling than just a few months back I’ll try to summarise how I see it.
From a strictly genetic point of view, one problem was that even CHG were already to “eastern” for a 2 way admixture with EHG to form Yamnaya. Always some amount of ANF and/or WHG was needed. Now these new Eneolithic_steppe samples lack that extra ANF/WHG admixture, so a straight EHG/CHG seems like it would work just fine. Something like Sarazm_Eneolithic seems to be again too “eastern”, though if you count that it’s not completely necessary that the admixture was just with EHG, but maybe something like Ukraine_Mesolithic or Neolithic too, that could theoretically work.
But this has several problems:
– These steppe Eneolithic samples don’t seem to show to be recently admixed. Certainly not with neolithic populations from the Caucasus, but even with Mesolithic (CHG) ones. The origin of this population looks temporarily and, more important, culturally, Mesolithic. So you’d need a very early migration to the steppe, at which point it becomes rather irrelevant from a linguistic and cultural point of view.
– Even if we prefer to assume that this population is recently admixed, we need some colourful scenario to explain the admixture event to match what we see from mtDNA and Y-DNA. We have no evidence of R1 in SC Asia so far, but plenty of it among Eastern European HGs. So we’d be forced again to accept a scenario where a CHG-like population migrated to the steppe (and if it was recent, they would be the Neolithic/Chalcolithic pastoralists), just to be absorbed by EHGs roaming around, who would borrow the culture and women, somehow marginalizing or somehow getting rid of the CHG-like males and their own EHG females. Not something very parsimonious. I think we can leave this kind of scenario behind.
So I not only see the possibility of a recent migration from SC Asia to form Yamnaya rather unlikely. More importantly, I think it doesn’t matter at all for any IE related question.
The possibilities of PIE being from SC Asia (be it India itself or somewhere else in what was later called Ariana), have a much better explanation through a southern route, related to Armenia_ChL, Maykop and the later influx of more specifically Iran_ChL kind of admixture that seems to have been the most important one in West Asia and the southern Balkans. It requires a complex scenario, but the spread of IE languages is a complex phenomenon, so that’s not a big problem. The problem is to match all the details in a coherent way. Something that will take time.
Really happy for your Blog.
I thought you would go dark. Really enjoy reading you.
Agreeing or disagreeing… all makes sense.
thanks.
@OM
Thanks! I hope that here we can all agree or disagree peacefully and respectfully. It’s probably going to be a more quiet place that others, but I hope it stays interesting, especially in the comments.
Alberto,
Perhaps you have misunderstood. What I m implying is – tbh a Johanna Nichols’ idea – that as IE groups expanded from around Bactria the Caspian Sea stood in the way and split the IE expansion two ways – one which went westward towards Iran, Caucasus and Anatolia and another which went North to reach the steppe and then this northern group also migrated westward on the steppe trajectory to reach Eastern Europe.
So what I m implying is that perhaps the Eneolithic steppe, steppe Maykop and Yamnaya represent this northern group which had split up from the Southern groups in Central Asia itself and again regained contact with the southern group in North Caucasus after having migrated from eastern steppe. So I m advocating that the Northern steppe group is another group of IE speakers that had turned nomadic while the southern group that formed Maykop and Armenia_Chl was a southern primarily agricultural group.
This may explain the Iran_N admixture without accompanying ANF in the steppe. As for Yamnaya having ANF and WHG, it could be as a result of an independent mixing with EEF group from the West coming from Europe.
Eventually as the southern and northern groups interacted again between North Caucasus and the steppe, the cultural tool kit was easily transferred to the steppe groups from Maykop without significant y-DNA influence.
In another words things may have been quite complex.
Having said so, it is undoubted that there is more solid evidence of interaction along the southern route between South Central Asia and the Near Eastern Chalcolithic. As this paper affirms, the groups ranging from Anatolian_Ch, Armenia_Ch, Iran_Ch to the North Caucasus were all part of a cultural interaction and I speculate that the main language or languages in this region of interaction was Indo-European. To me this is the biggest takeaway from this paper.
As for the R1b link, what is clear so far is that the Yamnaya R1b Z2103 is not found across Europe but is restricted in Eastern Europe is groups such as the Bashkirs but it is found across Anatolia, Armenia and all the way to South Asia with some Central Asian groups having a high proportion of it . All these Southern regions do not have any links with Yamnaya directly and the latter steppe_mlba groups are dominated with R1a. So how did R1a Z2103 spread across this vast Southern landscape ? It is an interesting question.
We may argue that this proves that R1b Z2103 spread from a Southern region and not from the steppe and this has merit because, besides the controversial R1b-Z2013 from Haji Firuz Chalcolithic , we also have R1b -M269 from Armenia_EBA and likely from the low coverage Darra I Kur from Afghanistan at 2600 BCE.
But what was the path ? There is no evidence of R1b expansion in Chalcolithic Caucasus – based on this current paper. Yet R1b – Z2103 shows founder effect not only in Yamnaya but also in some Central Asian groups..
@jaydeepsingh,
However the paper is about that trail and path. For the purpose of managing their samples, papers published and the way issues were looked at over recent and not so recent years, in this paper they isolate the Maykop issue and got done with.
Next is the publishing of the last piece of the puzzle.
Actually russian archaeology has seen that trail for long. They well describe cultural movements to which I needed to use google translator, from shulaveri to what they call pricked pearl culture and we mostly know as svobodnoe…
@jaydeep,
Bactria the Caspian Sea, or better said the guys that lived beyond Kopet Dag, moved to transcausia to be part of the movement that dislodged PIE.
https://r1b2westerneurope.blogs.sapo.pt/from-the-ubaid-and-kura-araxes-8426
@Jaydeep
Yes, I can mostly agree with that as a possibility. Though I still think that the northern route has the problems I stated above (steppe_eneolithic not looking as recently admixed between EHG and CHG, R1b being much ore likely EHG than CHG in such a mix, and mtDNA being CHG, plus invisible cultural influence in such group from a more developed SC Asian Chalcolithic population).
Most of modern R1b-Z2103+ south of the steppe is probably the result of Late or Post-Yamnaya movements across the Caucasus (and earlier to the Balkans). We do have evidence for this admixture in MLBA and IA samples from Armenia, NW Iran, EBA Balkans,… But if we do find a plausible source in SC Asia Eneolithic for this steppe population, then things would change. Maybe R1b-M269 came from that area, but right now it’s much less likely with the data we have. We still need more 5th mill. samples from several areas to know this with certainty.
Alberto,
That progress 2 R1b1 guy had a trepantion.
Since the oldest trepanation (apart from the france weird thing 6300bc) is a “shulaverians” at Chalaghantepe in Azerbaijan… who do you thing teach them to do it?
btw… you are rigth. They know what the subclades are. Obvious. But this paper was not the time to raise that issue.
Thanks for the blog, your comments have always been reasoned and reasonable,
Alberto,
I have a small request. Can you try modelling the steppe_Eneolithic and the Yamnaya using Sarazm_EN or Geoksiur_EN as one of the sources with other sources being Globular_Amphora & EHG ? Can you check if its a workable model ?
Regards,
Jaydeep
@Jaydeep
The Steppe_Eneolithic samples are still not available. I have tried before to see if the SC Asian samples can improve the models for Yamnaya, but on their own they don’t work well. They require something more “western” than EHG too, so I add Ukraine_Mesolithic too (Sarazm I4910 has some AASI, that’s why I add both samples separately):
Yamnaya_Samara
Sarazm_Eneolithic:I4290 40.4%
EHG 30.2%
Ukraine_Mesolithic 21.7%
Globular_Amphora 7.6%
Geoksiur_Eneolithic 0.1%
Ganj_Dareh_N 0%
Sarazm_Eneolithic:I4910 0%
Ukraine_N 0%
Distance 6.6525%
The distance of 6.6% is not good at all. Adding CHG too things improve:
Yamnaya_Samara
EHG 49.5%
CHG 28.2%
Sarazm_Eneolithic:I4290 10.3%
Globular_Amphora 6.3%
Ukraine_Mesolithic 5.7%
Ganj_Dareh_N 0%
Geoksiur_Eneolithic 0%
Sarazm_Eneolithic:I4910 0%
Ukraine_N 0%
Distance 5.5654%
Here another run I did a while back with all individuals:
https://docs.google.com/spreadsheets/d/1_rtImhfZrG4rSMKzPVulIKkA42_jT1o9meDGEameDUw/edit?usp=sharing
But notice that the models are still quite poor. And this is one of the things that always made me a bit unconvinced about Yamnaya being a mix of EHG and CHG from the Eneolithic. The models simply don’t work as good as they should. With D-stats you can also notice that CHG is ok, but not nearly as good as you would expect if these were the real populations that admixed in the steppe.
That’s one more reason why I think that Steppe_Eneolithic is a population that we could call “native” to the North Caucasus (it’s exact range still unknown, but probably extended to the North Caspian shores). It looks Mesolithic or older to me, though not enough tests with these samples have been performed yet to really know.
Alberto,
Or anybody for that matter.
Sarazm_Eneolithic are really different one from the other. Any thoughts on why?
Alberto,
Thanks a lot. I looked at the Khwalynsk samples in the recent Narasimhan et al paper and it is quite interesting. Looking at figures S3.22 & S3.23, in terms of allele sharing these steppe_Eneolithic samples are significantly much closer to WSHG than most steppe populations and this includes the Okunevo samples which are modelled as essentially 55 % WSHG with the rest being Han & Iran_N. Looking at f3 stats, the highest scores for steppe_Eneolithic seem to come when the steppe source in AG3 followed by WSHG. EHG is not in the picture. Looking at the distal model for Khwalynsk (table S3.45), though it is modelled as EHG + Iran_N, it is a very poor fit with p value being 0.058. So there is no certainty in my opinion that the steppe ancestry in Steppe_Eneolithic is necessarily from EHG. It looks closer to AG3 or WSHG.
For the Yamnaya or steppe_EMBA, the Narasimhan et al paper models them as Khwalynsk_EN + Haji_Firuz_C, while the Wong et al paper models them as EHG + CHG + ANF/EEF. Essentially, also keeping in mind earlier attempts, it seems to boil down to 1 steppe component (EHG or something related), one ANF/EEF component & one Iran_N/CHG component.
It is interesting that in your model, Geoksiur_EN is preferred while Seh_Gabi & Haji_Firuz are not. And while Dali_EBA is preferred, WSHG is not. What according to you is the best modelling of Yamnaya ?
Regards.
Olympus Mons,
As Alberto said, one sample has AASI while the other does not. Sarazm is a site which has clear links with chalcolithic cultures south of the Hindu Kush. Have a look,
https://www.researchgate.net/publication/308795205_2014-Contacts_across_the_Hindu_Kush_in_the_Early_Bronze_Age_Additional_Insights_from_Sarazm_-_Soundings_11-11A_Tajikistan
Jaydeep,
Tks. Will read it.
@Jaydeep
Yes, Khvalynsk being very EHG is normal that it shares more alleles with WSHG than other steppe groups. I don’t think it means it has any direct input from WSHG or AG3, though. I think that Khvalynsk is just an early migration north from the Steppe_Eneolithic population sampled in the North Caucasus, where a few of them went up the Volga and settled in the Samara region, where they got a good amount of EHG admixture before disappearing.
Yamnaya for me descends from that same Steppe_Eneolithic population from the North Caucasus steppe, only with some 10% “European” admixture from around Ukraine (probably Ukraine_N + Tripolye or some similar combination).
@OM
Yes, like Jaydeep said, the 2 Sarazm samples are a bit different in that one has a small amount of AASI that the other lacks. That’s the only difference I can see so far.
If I am not mistaken, the curve of the f3 statistics for the sample tested has a parabolic behavior, reaching its minimum when the contrasted population is at the midpoint between the allele frequencies of the reference pops. This implies that the largest yamna signal in f3 could result from a more “centered” position than eneolithic steppe with respect to the two poles of f3, EHG and CHG. In the PCA (fig.2-d) I did not observe a distinguishable displacement of yamna with respect to eneolithic steppe towards the chg/Iran cluster, without appreciating displacement in PC1, but rather towards the “south” in PC2. If we add the signal increase in F3, ¿could a subtle contribution of neolithic anatolia(who carries whg) or CHG (or both) be the cause of that signal increase in f3, centering yamna regarding a position, i belive, closer to EHG occupied by eneolithic steppe?
That´s said in “Genome-wide patterns of selection in 230 ancient Eurasians”, Admixture between populations of Near Eastern ancestry and the EHG began as early as the Eneolithic (5200-4000 BCE), with some individuals resembling EHG and some resembling Yamnaya”, this are f-statistics in the paper.
EHG Yamnaya_Samara Armenian Chimp −0.00191 −6.1
EHG Yamnaya_Kalmykia Armenian Chimp −0.00180 −5.4
Samara_Eneolithic Yamnaya_Samara Armenian Chimp −0.00100 −3.3
EHG Poltavka Armenian Chimp −0.00175 −4.9
the lowest values correspond to Samara_Eneolithic, vs. EGH, so we can limit the period of appearance of the, in study terminology, “Near Eastern ancestry”(Armenian which carries CHG) from the last EHG onwards(maybe 5500 bce+-, even maybe before).
In the same way, “Early Neolithic genomes from the eastern Fertile Crescent” D-stats(Neo_Iranian, Pre_neolithic, Ancient, Khomani) (Neo Iranian is Abdul Hosein 1&2, but works in similar way whith the others)
AH1 CHG Samara_Eneolithic Khomani -0,0203 -3,041
AH1 CHG Yamnaya_Kalmykia Khomani -0,033 -6,41
AH1 CHG Yamnaya_Samara Khomani -0,0357 -7,658
AH1 EHG Samara_Eneolithic Khomani -0,1317 -21,073
AH1 EHG Yamnaya_Kalmykia Khomani -0,0657 -13,449
AH1 EHG Yamnaya_Samara Khomani -0,0811 -17,714
AH2 CHG Samara_Eneolithic Khomani -0,044 -5,7
AH2 CHG Yamnaya_Kalmykia Khomani -0,0468 -8,565
AH2 CHG Yamnaya_Samara Khomani -0,0503 -9,707
AH2 EHG Samara_Eneolithic Khomani -0,1543 -23,06
AH2 EHG Yamnaya_Kalmykia Khomani -0,0782 -15,445
AH2 EHG Yamnaya_Samara Khomani -0,0918 -19,589
detect the presence of Neolithic Iranian in Samara Eneolithic, substantially increased in yamnaya (increase of relationship with chg to the same extent that dilutes ehg), so the signal can be interpreted, I think, not as a greater presence in yamna of ehg but of chg/iran_neol.
@Alberto
thank you for the reference to the practice of trepanacion, but poking into bibliographies of the network seem quite widespread practices, perhaps not discriminant in this case.
@Anthro Survey
What do you mean with “Note that Caucasus and Eneolithic Steppe demes(as defined by paper) take different CHG-related nodes”, if I understand correctly Fig. 5. Admixture Graph, they both take from Basal Eurasian at the same level (zero steps).
@Jaydeepsinh Rathod
good points
In Caucasus paper, supplementary Fig. 5 results of D-statistics D (EHG, eneolithic steppe, X, Mbuti) are negative and significant from Anatolia Tepecik Ciftlik to CHG, through Ganj Darej iran neolithic.
Regarding the neolithic Tepecik Ciftlik, it shares a chg/iran_neolitic component with Peloponnese Neolithic that indicates an early presence in east mediterranean, and that does not seem to be present in other samples of the Anatolian Neolithic(e.g.Barcin-Mentese) or European. Later it is present in Armenia/Anatolian_Calcolithic/BA and in Minoan/Mycenean (more in the latter).
The genomic history of southeastern Europe
“these ‘Peloponnese Neolithic’ individuals, dated to around 4000 bc, are shifted away from WHG, and towards CHG, relative to northwestern-Anatolian Neolithic and Balkan Neolithic individuals”
The Demographic Development of the First Farmers in Anatolia
“D statistics results revealed genetic affinity between Caucasus hunter-gatherers (CHGs) and one of the individuals from Tepecik-Ciftlik, Tep003, which was greater than the rest of the individuals from Tepecik-Ciftlik and other Neolithic individuals”. Although observing the PCA we see that this affinity can be extended to the rest of Tepecik-Ciftlik samples, “Tepecik-Ciftlik individuals [..] were positioned at a peripheral position within the whole cluster and displayed high within-group diversity. Pairwise f3 statistics between populations also showed significant differentiation between Boncuklu and Tepecik-Ciftlik populations”.
It seems to point the formation of a complex of relations in the Neolithic, in the area between Anatolia and Iran, precursor of cultures such as Kura-Araxes and Maykop.
In summary, i think that the possibility, in my opinion probably and chronologically possible, of a neolithic influx, hurrying mesolithic, coming originally from Iran (the route is belatedly witnessed in Namagaza_CA) that flows into the steppe to form the “steppe ancestry”, should be considered. I could not say, however, if that chg/iran component in the east of the Mediterranean comes from iran/caucasus or from a third source “Basal Eurasian”, although in the case of the formation of the “steppe ancestry” I would choose the former. On the other hand, it may have been possible that, also in small doses, the Anatolian Neolithic reached the “eneolithic steppe” by another route, different from that of Caucasus, as is seen in Ukraine_N_Outlier.
found out about this channel recently. A welcome change from that other one!
“Reich lab” treated the Central Asian samples with extreme prejudice, derision, and dismissiveness. They carelessly typed the samples using an untested new computerized sequencer and they threw away samples by the hundred, for “contamination” and “illogical” results. I’d wager much of this was likely R1a1 and R1b. I don’t blame Reich, but he”s surrounded by supremacists of a certain ilk.
@darklord
How do you know this? Is it elaborated in the paper?
If the screening is unbiased and only dependent on sample quality, then it should still be representative
Back from the holidays. I’ll try to catch up in the next few days with some posts, if time permits.
@Postneo, thanks.
@Algan mardi
Re: the f3 stats, I’m not sure about the parabolic behaviour that you mention. Maybe that’s correct when everything else is equal. But usually there are other factors that are not equal and have bigger influence. For example, post-admixture drift.
The Eneolithic_steppe samples are from around 4300 BCE, and they are already around 50/50 EHG/CHG, so they are in the middle of the curve. And being much closer to the putative admixture event, the magnitude of the stats should be quite bigger than in Yamnaya. This can better be explained as those samples not being recently admixed. Yamnaya, on the other hand, had more recent admixture from different sources, which increases the magnitude of the stats.
And as I mentioned above in a reply to Jaydeep, there’s still the problem of the Y-DNA and mtDNA if you want to go for a recent admixture of two different populations (EHG-like and CHG-like). Plus the archaeological vacuum to link the more advanced Iranian/Caucasian cultures to the Steppe_Eneolithic ones from the North Caucasus steppe.
So overall, I’m quite sceptic about that scenario. Though the cultural impact from Iran/Caucasus in the later steppe cultures is very important, though (more on this soon).
@darkpath, I don’t see any evidence for that. The latest papers have clearly eroded the simplistic EHG = PIE equation, so why would we think that they have hidden R1 samples from outside the steppe? What the ancient DNA data available today shows is a very weak correlation between R1 and IE languages, so the origin of R1 (most likely in Eastern Europe) is becoming increasingly irrelevant for the IE question. Rather than some conspiracy, there might be some unconsciously (?) biased interpretations of the data in Narashimhan et al. 2018 preprint (more on that soon too).
@Alberto
The scenario is presented in speculative terms and, for now, limitedly testable.
As I said, I’m not sure about that parabolic behavior in f3 either, which if it seems correct is to interpret a negative value as probable mix and a positive one as absence of proof, because the drift that quotes can happen, in the case of f3-Outgroup would measure the shared drift.
Nor would I be sure that a closer proximity of the population contrasted at the time of the mixture gave higher values in f3, ¿could it depend, not only on the moment but also on its initial position with respect to the reference poles?
Certainly, Eneolithic steppe looks almost 50% EHG/CHG; in supplemental information, “Eneolithic steppe individuals deriving more than 60% of ancestry from EHG and the remainder from a CHG related basal lineage(it catches my attention that they use the expression basal)”. This seems supported in Broushaki et al, both Yamna and Samara eneolithic choose to EHG, but faced with CHG emerges the cluster of this along with Iran N, which seems to make yamna/steppe more EHG than CHG.
I recognize, in any case, that even without knowing how to affect the results of confronting the population tested with ‘the same cluster’, in the D-stats, Samara_Eneolithic&Yamnaya still prefer CHG rather than Iran neolithic, so as you say, it could be that the impact, was mostly cultural.
d z
Samara_Eneolithic AH2 CHG Khomani -0,1353 -19,217
Samara_Eneolithic AH4 CHG Khomani -0,1212 -18,034
Yamnaya_Samara AH2 CHG Khomani -0,1163 -21,171
Yamnaya_Kalmykia AH2 CHG Khomani -0,1159 -20,049
Samara_Eneolithic AH1 CHG Khomani -0,1077 -16,554
Yamnaya_Samara AH4 CHG Khomani -0,1042 -19,678
Yamnaya_Kalmykia AH4 CHG Khomani -0,1023 -19,088
Yamnaya_Samara AH1 CHG Khomani -0,0881 -16,716
Yamnaya_Kalmykia AH1 CHG Khomani -0,0866 -15,641
Samara_Eneolithic WC1 CHG Khomani -0,0743 -13,842
Yamnaya_Samara WC1 CHG Khomani -0,0585 -13,207
Yamnaya_Kalmykia WC1 CHG Khomani -0,0558 -11,965
Yamnaya_Kalmykia AH1 EHG Khomani 0,0946 18,433
Yamnaya_Kalmykia AH4 EHG Khomani 0,0998 18,365
Yamnaya_Kalmykia AH2 EHG Khomani 0,1006 17,959
Yamnaya_Samara AH1 EHG Khomani 0,1032 21,582
Yamnaya_Kalmykia WC1 EHG Khomani 0,1034 24,006
Yamnaya_Samara AH4 EHG Khomani 0,1064 20,103
Yamnaya_Samara AH2 EHG Khomani 0,1097 20,464
Yamnaya_Samara WC1 EHG Khomani 0,111 26,586
Samara_Eneolithic AH1 EHG Khomani 0,1498 24,488
Samara_Eneolithic AH4 EHG Khomani 0,1563 24,204
Samara_Eneolithic WC1 EHG Khomani 0,1567 30,598
Samara_Eneolithic AH2 EHG Khomani 0,1585 22,921
On the other hand, you are right that the Y-DNA and mtDNA problem would persist in recent admixture of two different populations. However, without having only a strictly temporary view, it would not be a priori disposable, in my opinion, to consider the origin of the r1b in Yamna, ultimately, Iran (or Central-South Asia). See the Balanovsky R1b iron iran sample F38, which although understood as a derivative of yamna (I have not yet read the paper of central Asia formation), it could also be considered as derived from an ancestral source common to both and originally from Iran area. In its support, the diversity of r1b-269 indicated in “Ancient Migratory Events in the Middle East New Clues from the Y-Chromosome Variation of Modern Iranian” Table S7, with maximum values of age based on the diversity of microsatellites for Iran and Turkey (to be taken into account also for R1a). To the detriment, AFAIK, the absence of r1b samples in meso/neolithic Iran, although there is a J in Karelia EHG, and Broushaki “our male Iron Age genome from Tepe Hasanlu in NW-Iran shares greatest similarity with Kumtepe6 even when compared to Neolithic Iranians. We inferred additional non-Iranian or non-Anatolian ancestry in F38 from sources such as European Neolithics and even post-Neolithic Steppe populations. Consistent with this, F38 carried a N1a sub-clade mtDNA, which is common in early European and NW-Anatolian farmers. In contrast, his Y-chromosome belongs to sub-haplogroup R1b1a2a2, also found in five Yamnaya individuals”.
A laps with the Dstats, would seem to support the latter, F38 prefers EHG, Yamna and Samara eneolithic against the Iranian Neolithic and discards CHG that form their own cluster with Iran_N.
However, it does not fail to mention that “Kumtepe6, a ~ 6,750 year old genome from NW-Anatolia, was more similar to Neolithic Iranians than any other non-Iranian ancient genome […] These patterns indicate that post-Neolithic homogenization in SW-Asia involved substantial bidirectional gene flow between the East and West of the region, as well as possible gene flow from the Steppe”, leaving the door open to the complex neolithic relationship between anatolia/iran that could be an origin of the CHG component in Eneolitic steppe.
d z
AH2 F38 Samara_Eneolithic Khomani -0,0523 -5,827
WC1 F38 Samara_Eneolithic Khomani -0,047 -7,433
AH2 F38 Yamnaya_Samara Khomani -0,0382 -6,253
WC1 F38 EHG Khomani -0,0382 -6,079
WC1 F38 Yamnaya_Kalmykia Khomani -0,0365 -7,097
WC1 F38 Yamnaya_Samara Khomani -0,0359 -7,571
AH4 F38 Samara_Eneolithic Khomani -0,0353 -4,221
AH2 F38 EHG Khomani -0,0334 -4,061
AH2 F38 Yamnaya_Kalmykia Khomani -0,0327 -4,958
AH4 F38 EHG Khomani -0,0327 -4,274
AH1 F38 EHG Khomani -0,0267 -3,61
AH1 F38 Samara_Eneolithic Khomani -0,0254 -3,129
AH1 F38 Yamnaya_Samara Khomani -0,0247 -4,238
AH4 F38 Yamnaya_Samara Khomani -0,0238 -4,203
AH1 F38 Yamnaya_Kalmykia Khomani -0,0146 -2,361
AH4 F38 Yamnaya_Kalmykia Khomani -0,0128 -2,108
WC1 F38 CHG Khomani -0,0092 -1,435
AH1 F38 CHG Khomani 0,0175 2,337
AH4 F38 CHG Khomani 0,0301 3,997
AH2 F38 CHG Khomani 0,0491 5,799